



Design a primer to linearize plasmid backbone
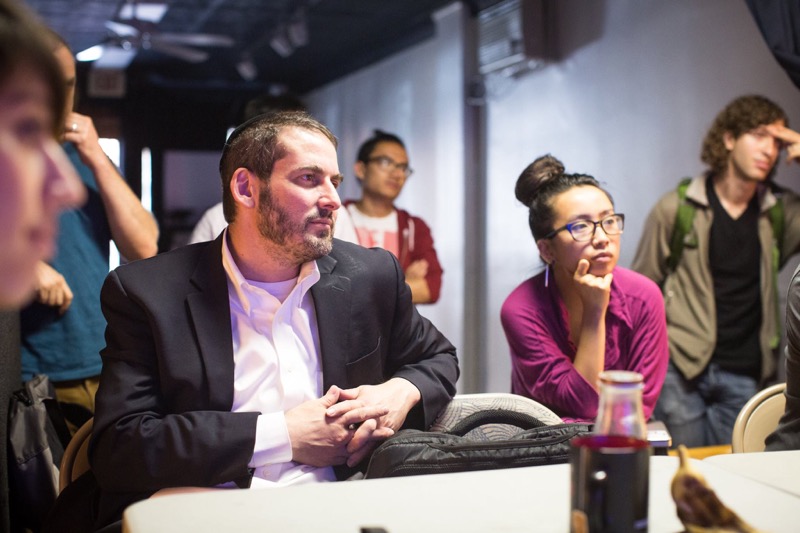
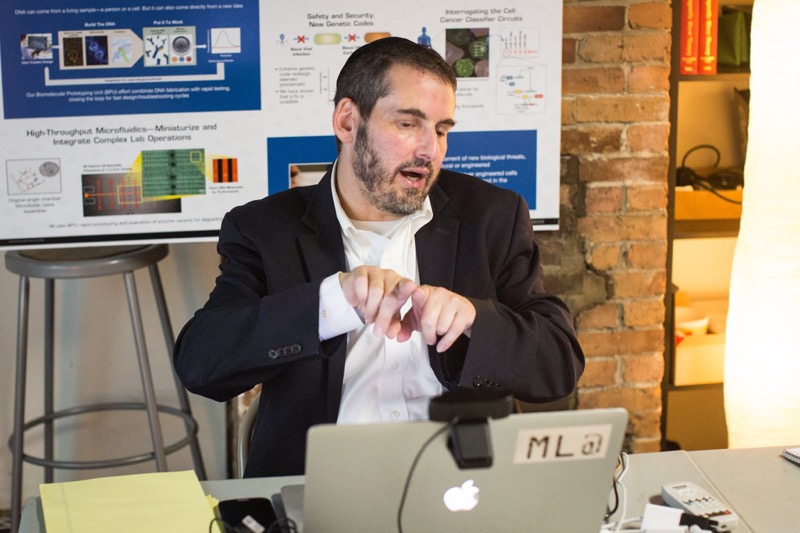
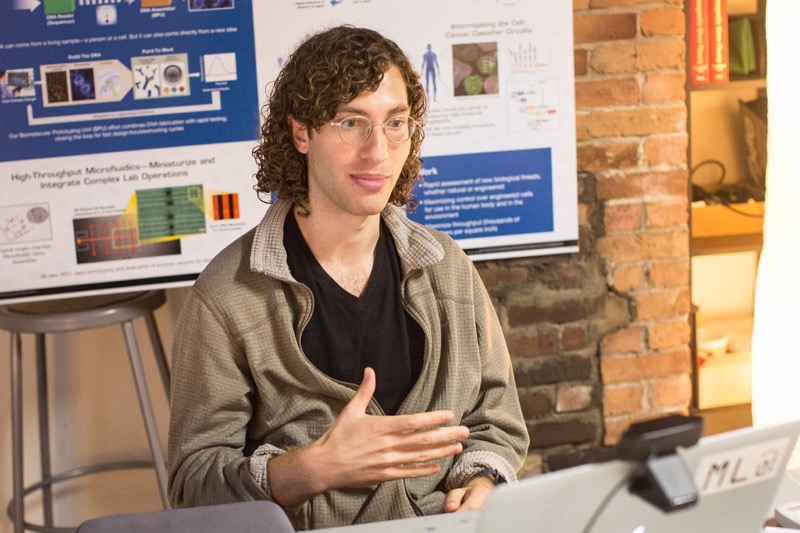
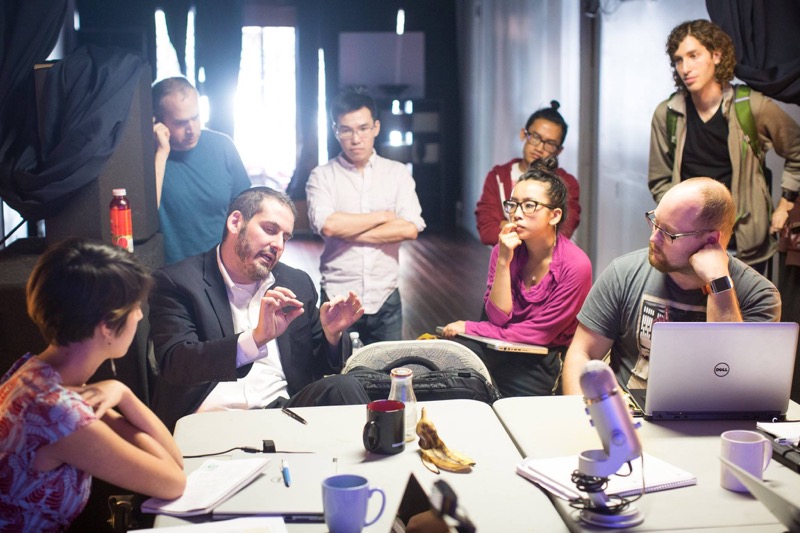
Theory: Use NuPack to help select 18 bp priming sites that amplify a ~2.25 kb region of pUC19 (NEB) immediately upstream of the plac promoter and downstream of the start of lacZalpha. The resulting amplicon excludes the plac promoter and n-terminus of lacZalpha, which enables you to swap in a gene of interest under the control of a promoter of interest using Gibson Assembly later on in your workflow. Design one pair of oligos that prime optimally and another that prime poorly. Describe the PCR thermocycling program that uses Phusion polymerase (NEB) for these pairs. Crucially, determine annealing temperatures and extension times.![]()
PRIMER 1
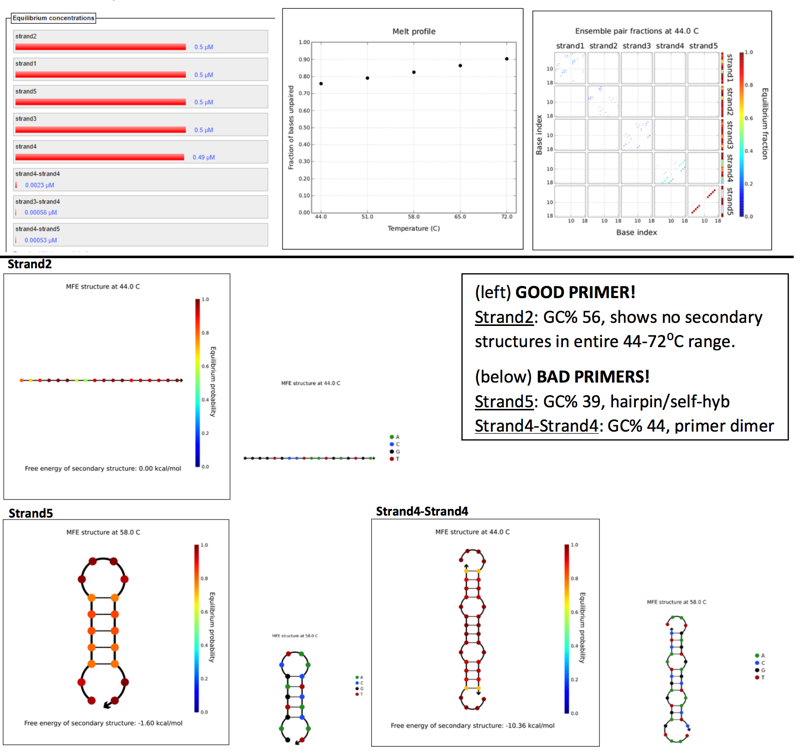
Scanned 18bp segments, at 4nt step resolution for 5 strand species.Parameters set by Phusion polymerase, at primer concentration of 0.5μM and several annealing temps ranging between 45-72⁰C (5 temps, with 7⁰C increments), assuming TA is 3-5⁰C below TM.
Results: (left to right) Equilibruium concentrations histogram, Melt profile (fractio of bases unpaired), and Ensemble pair fractions.
PRIMER 2
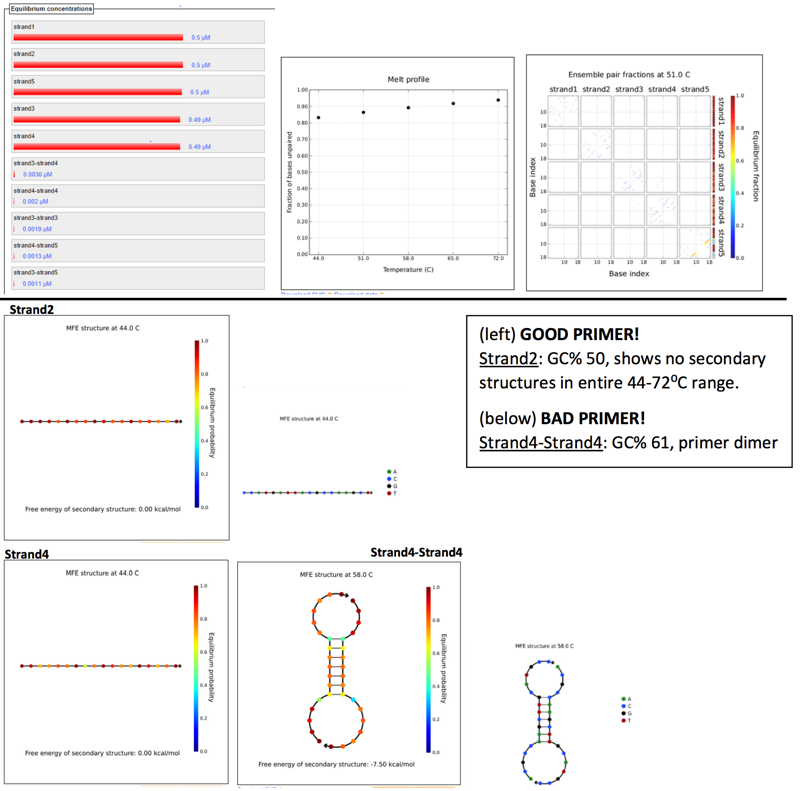
Scanned 18bp segments, 4nt step res for 5 strand species. (Reverse complement to original sequence).Parameters set by Phusion polymerase, at primer concentration of 0.5µM and several annealing temps ranging between 45-72⁰C (5 temps, with 7⁰C increments), assuming TA is 3-5⁰C below TM.
Results: (left to right) Equilibruium concentrations histogram, Melt profile, and Ensemble pair fractions.
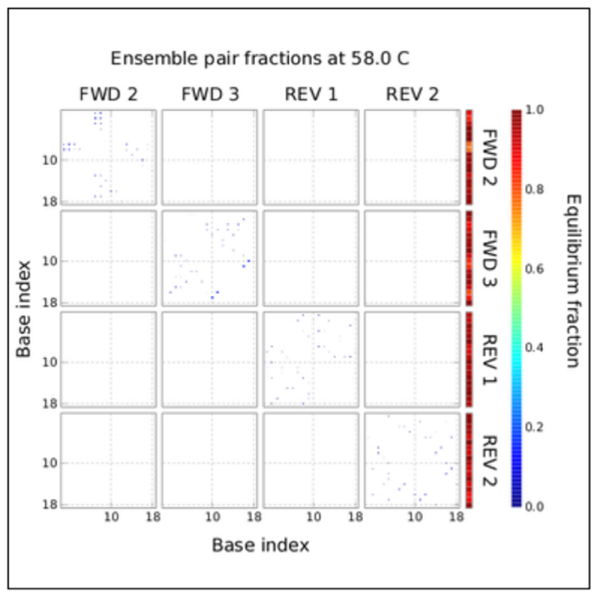
Good PRIMER Set
Similar GC%, and melting temps. No instances of self-annealing or primer dimers between them. Produces a 2.6Kbp amplicon (a little on the large side, target 2.3Kbp)
PCR @ Annealing Temp: 55⁰C PCR @Extension Time (@ Phusion 15 sec/kb recommendation): 40 sec
BAD PRIMER SET:
Both primers have GC% out of the 40-60% range, and respectively too low and too high. Because we have to choose a single annealing temperature to use in PCR cycles, the wide range between their GC%/temps adds to the inefficacy of these primers.
![How to grow [almost] anything class Viirj Kan](img/logo.png)